One-way ANOVA Tests in R
Here, we discuss the one-way analysis of variance (ANOVA) test in R with interpretations, including, f-value, sum of squares, mean squares, p-values, and critical values.
The one-way ANOVA test in R can be performed with the
anova() or oneway.test() function from the
base "stats" package.
The one-way ANOVA test can be used to test whether the means of the populations where two or more independent random samples come from are equal (as stated in the null hypothesis) or not. With the typical equal variance assumption, it is an extension of the pooled two sample t-test from two means (or groups) to three or more means.
In the one-way ANOVA test, the test statistic follows an F-distribution when the null hypothesis is true.
| Question | Are the group means equal? |
| Null Hypothesis, \(H_0\) | \(\mu_1 = \mu_2 = \dots = \mu_G\) |
| Alternate Hypothesis, \(H_1\) | At least one group’s mean is different from the rest. |
Sample Steps to Run a One-way ANOVA Test:
# Create the group data for the one-way ANOVA test
# Each row contains values for one group
y = c(7.2, 5.4, 5.2, 1.4,
8.4, 4.8, 6.5,
5.8, 6.2, 8.4, 4.8, 6.5)
groups = c("A", "A", "A", "A",
"B", "B", "B",
"C", "C", "C", "C", "C")
# Run the one-way ANOVA test with specifications
anova(lm(y ~ groups))Analysis of Variance Table
Response: y
Df Sum Sq Mean Sq F value Pr(>F)
groups 2 7.138 3.5690 1.0269 0.3965
Residuals 9 31.279 3.4754
One-way analysis of means
data: y and groups
F = 1.0269, num df = 2, denom df = 9, p-value = 0.3965Argument (anova()) |
Argument (oneway.test()) |
Usage |
| lm(y ~ x) | y ~ x | y contains the sample data values, x specifies the group they belong. |
| var.equal | Set to TRUE for equivalence with
anova().FALSE is unequal variance
assumption, an extension of Welch’s t-test. |
Creating a One-way ANOVA Test Object:
# Create data
y = rnorm(30)
groups = rep(c("A", "B", "C"), each = 10)
# Create object
aov_object = anova(lm(y ~ groups))
# or
owt_object = oneway.test(y~groups, var.equal = TRUE)
# Extract a component
aov_object$`F value`[1][1] 1.17051 F
1.17051 anova() Component |
oneway.test() Component |
Usage |
| aov_object$`F value`[1] | owt_object$statistic | Test-statistic value |
| aov_object$`Pr(>F)`[1] | owt_object$p.value | P-value |
| aov_object$Df | owt_object$parameter | Degrees of freedom |
| aov_object$`Sum Sq` | Sum of squares | |
| aov_object$`Mean Sq` | Mean squares |
1 Test Statistic for One-way ANOVA Test in R
The one-way analysis of variance test has test statistics, \(F\), of the form:
\[\begin{align*} F & = \frac{\tt{variance \; between \; groups}}{\tt{variance \; within \; groups}} = \frac{MS_{groups}}{MS_{residuals}} \\ & = \frac{\left[ \sum_{j=1}^G N_j(\bar y_{j\cdot}-\bar y_{\cdot \cdot})^2 \right] / (G-1)}{\left[ \sum_{j=1}^G \sum_{k=1}^{N_j} (y_{jk}-\bar y_{j\cdot})^2 \right]/ (N-G)}\\ & = \frac{\left[ \sum_{j=1}^G N_j(\bar y_{j.}-\bar y_{\cdot \cdot})^2 \right]/ (G-1)}{\left[ \sum_{j=1}^G (N_j-1) s_j^2 \right]/ (N-G)}. \end{align*}\]
For independent random samples that come from normal distributions, \(F\) is said to follow the F-distribution \(\left(F_{G-1,N-G}\right)\) when the null hypothesis is true, with \(G - 1\) numerator degrees of freedom, and \(N-G\) denominator degrees of freedom.
|
Source of Variation |
Degrees of Freedom (DF) |
Sums of Squares (SS) |
Mean Square (MS) |
F | P-value |
|---|---|---|---|---|---|
| Groups | \(G-1\) | \(\sum_{j=1}^G N_j(\bar y_{j\cdot}-\bar y_{\cdot \cdot})^2\) | \(\frac{SS_{groups}}{DF_{groups}}\) | \(\frac{MS_{groups}}{MS_{residuals}}\) | \(P(F_{G-1,N-G}>F)\) |
| Residuals | \(N-G\) | \(\sum_{j=1}^G \sum_{k=1}^{N_j} (y_{jk}-\bar y_{j\cdot})^2\) | \(\frac{SS_{residuals}}{DF_{residuals}}\) | ||
| Total | \(N-1\) | \(\sum_{j=1}^G \sum_{k=1}^{N_j} (y_{jk}-\bar y_{\cdot \cdot})^2\) |
\(G\) is the total number of groups,
\(N_j\) is the number of observations from group \(j\),
\(N\) is the total number of observations, \(\sum_{j=1}^G N_j\),
\(y_{jk}\) is the \(k\)th observation from group \(j\),
\(\bar y_{j\cdot}\) is the sample mean of group \(j\),
\(s_j^2\) is the sample variance of group \(j\),
\(\bar y_{\cdot \cdot}\) is the overall sample mean.
See also the two-way ANOVA test.
For a non-parametric test, see the Kruskal-Wallis test.
2 Simple One-way ANOVA Test in R
Enter the data by hand.
y = c(18.7, 20.3, 21.0, 18.2,
21.4, 21.1, 22.9, 19.5, 19.5, 18.5,
13.9, 23.7, 20.7, 21.9, 17.1)
groups = c("A", "A", "A", "A",
"B", "B", "B", "B", "B", "B",
"C", "C", "C", "C", "C")Check variability between and within groups with a boxplot.
The median lines appear to be on the same level, hence, there appears to be no group effects.
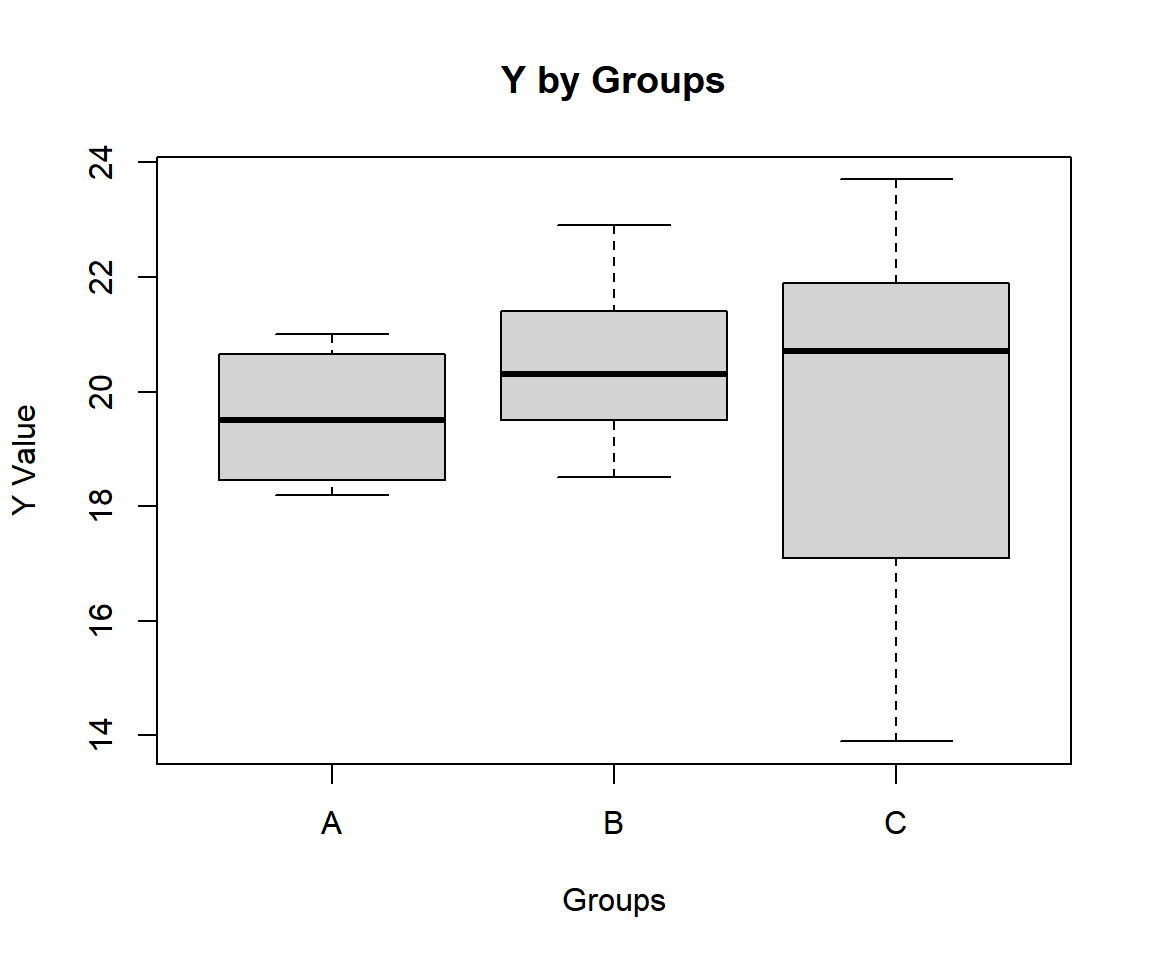
Simple One-way ANOVA Test Box Plot in R
The group and overall means, variances and lengths are:
A B C
19.55000 20.48333 19.46000 19.89333 A B C
1.736667 2.585667 15.488000 5.970667 A B C
4 6 5 15 For the following null hypothesis \(H_0\), and alternative hypothesis \(H_1\), with the level of significance \(\alpha=0.05\).
\(H_0:\) the group population means are equal (\(\mu_A = \mu_B = \mu_C\)).
\(H_1:\) at least one group’s mean is different from the rest.
Analysis of Variance Table
Response: y
Df Sum Sq Mean Sq F value Pr(>F)
groups 2 3.499 1.7495 0.2621 0.7737
Residuals 12 80.090 6.6742 Or:
One-way analysis of means
data: y and groups
F = 0.26213, num df = 2, denom df = 12, p-value = 0.7737The test statistic, \(F\), is 0.26213,
the degrees of freedom, are numerator df \(G-1= 2\), denominator df \(N-G= 12\),
the p-value, \(p\), is 0.7737.
Interpretation:
P-value: With the p-value (\(p = 0.7737\)) being greater than the level of significance 0.05, we fail to reject the null hypothesis that the group population means are equal.
\(F\) T-statistic: With test statistics value (\(F_{2, 12} = 0.26213\)) being less than the critical value, \(F_{2, 12, \alpha}=\text{qf(0.95, 2, 12)}\)\(=3.8852938\) (or not in the shaded region), we fail to reject the null hypothesis that the group population means are equal.
x = seq(0.01, 5, 1/1000); y = df(x, df1=2, df2=12)
plot(x, y, type = "l",
xlim = c(0, 5), ylim = c(-0.06, min(max(y), 1)),
main = "One-way ANOVA Test
Shaded Region for Simple Test",
xlab = "x", ylab = "Density",
lwd = 2, col = "blue")
abline(h=0)
# Add shaded region and legend
point = qf(0.95, 2, 12)
polygon(x = c(x[x >= point], 5, point),
y = c(y[x >= point], 0, 0),
col = "blue")
legend("topright", c("Area = 0.05"),
fill = c("blue"), inset = 0.01)
# Add critical value and F-value
arrows(2.5, 0.4, 0.26213, 0)
text(2.5, 0.45, "F = 0.26213")
text(3.885294, -0.04, expression(F[alpha]==3.885294))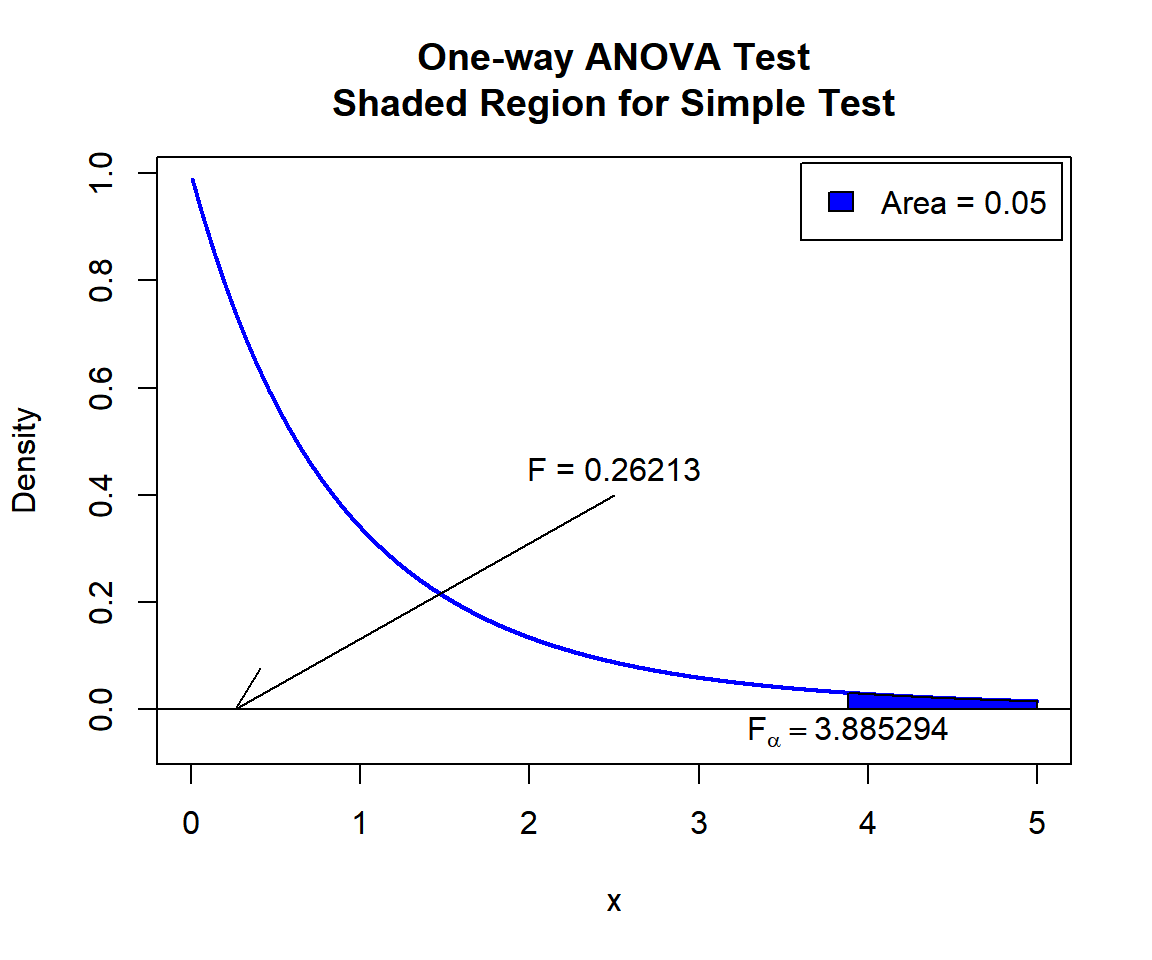
One-way ANOVA Test Shaded Region for Simple Test in R
See line charts, shading areas under a curve, lines & arrows on plots, mathematical expressions on plots, and legends on plots for more details on making the plot above.
3 One-way ANOVA Test Critical Value in R
To get the critical value for a one-way ANOVA test in R, you can use
the qf() function for F-distribution to derive the quantile
associated with the given level of significance value \(\alpha\).
The critical value is qf(\(1-\alpha\), df1, df2).
Example:
For \(\alpha = 0.1\), \(\text{df1} = 3\), and \(\text{df2} = 18\).
[1] 2.4160054 One-way ANOVA Test in R
Using the PlantGrowth data from the "datasets" package with 10 sample rows from 30 rows below:
weight group
1 4.17 ctrl
4 6.11 ctrl
5 4.50 ctrl
7 5.17 ctrl
12 4.17 trt1
17 6.03 trt1
19 4.32 trt1
20 4.69 trt1
23 5.54 trt2
30 5.26 trt2Check variability between and within groups with a boxplot.
boxplot(weight ~ group, data = PlantGrowth,
main = "Y by Groups",
xlab = "Groups",
ylab = "Y Value")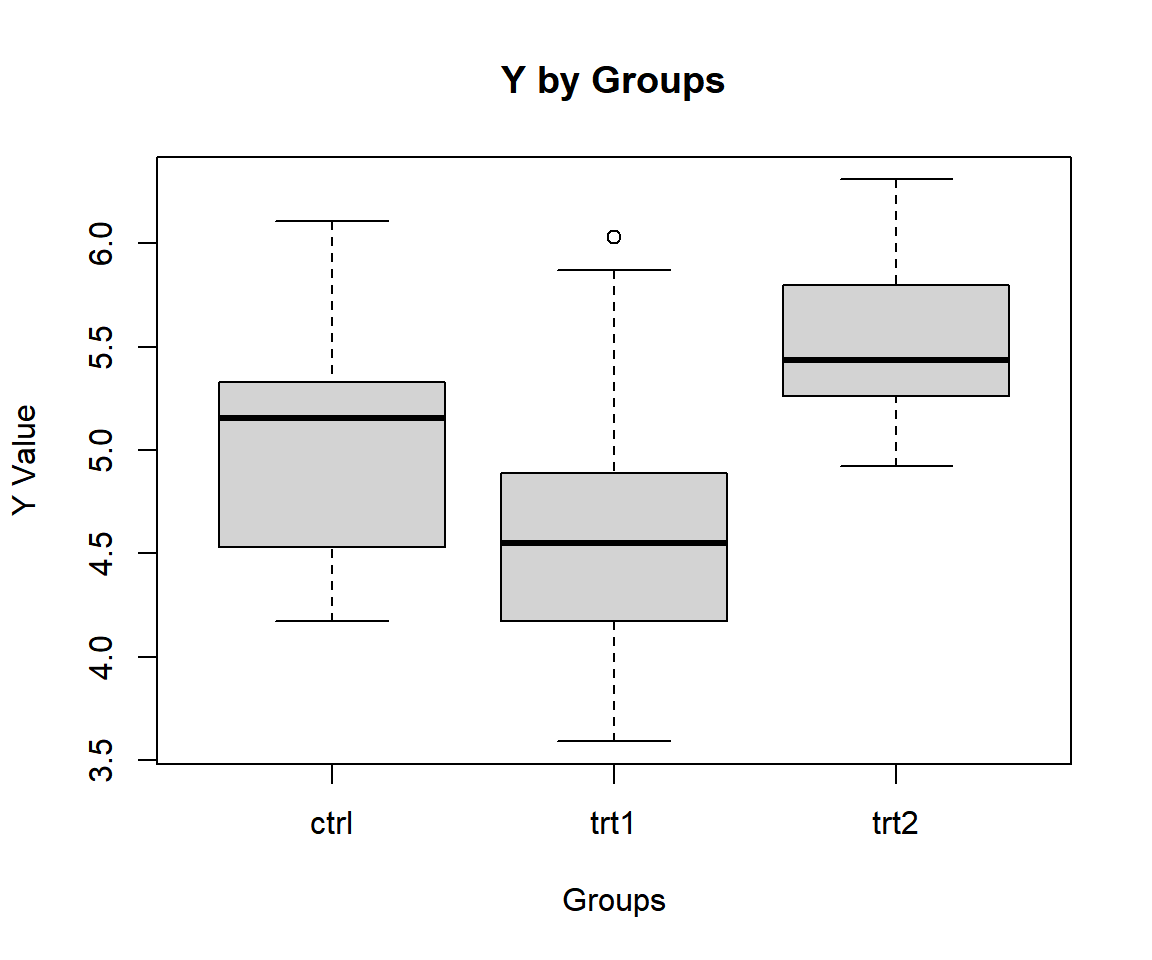
One-way ANOVA Test Box Plot in R
The group and overall means, variances and lengths are:
ctrl trt1 trt2
5.032 4.661 5.526 5.073 ctrl trt1 trt2
0.3399956 0.6299211 0.1958711 0.4916700 ctrl trt1 trt2
10 10 10 30 For the following null hypothesis \(H_0\), and alternative hypothesis \(H_1\), with the level of significance \(\alpha=0.1\).
\(H_0:\) the group population means are equal (\(\mu_{ctrl} = \mu_{trt1} = \mu_{trt2}\)).
\(H_1:\) at least one group’s mean is different from the rest.
Analysis of Variance Table
Response: weight
Df Sum Sq Mean Sq F value Pr(>F)
group 2 3.7663 1.8832 4.8461 0.01591 *
Residuals 27 10.4921 0.3886
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Or:
One-way analysis of means
data: weight and group
F = 4.8461, num df = 2, denom df = 27, p-value = 0.01591Interpretation:
P-value: With the p-value (\(p = 0.01591\)) being less than the level of significance 0.1, we reject the null hypothesis that the group population means are equal.
\(F\) T-statistic: With test statistics value (\(F_{2, 27} = 4.8461\)) being in the critical region (shaded area), that is, \(F_{2, 27} = 4.8461\) greater than \(F_{2, 27, \alpha}=\text{qf(0.9, 2, 27)}\)\(=2.5106087\), we reject the null hypothesis that the group population means are equal.
x = seq(0.01, 5, 1/1000); y = df(x, df1=2, df2=27)
plot(x, y, type = "l",
xlim = c(0, 5), ylim = c(-0.06, max(y)),
main = "One-way ANOVA Test
Shaded Region",
xlab = "x", ylab = "Density",
lwd = 2, col = "blue")
abline(h=0)
# Add shaded region and legend
point = qf(0.9, 2, 27)
polygon(x = c(x[x >= point], 5, point),
y = c(y[x >= point], 0, 0),
col = "blue")
legend("topright", c("Area = 0.1"),
fill = c("blue"), inset = 0.01)
# Add critical value and F-value
arrows(4, 0.3, 4.8461, 0)
text(4, 0.35, expression(F==4.8461))
text(2.510609, -0.04, expression(F[alpha]==2.510609))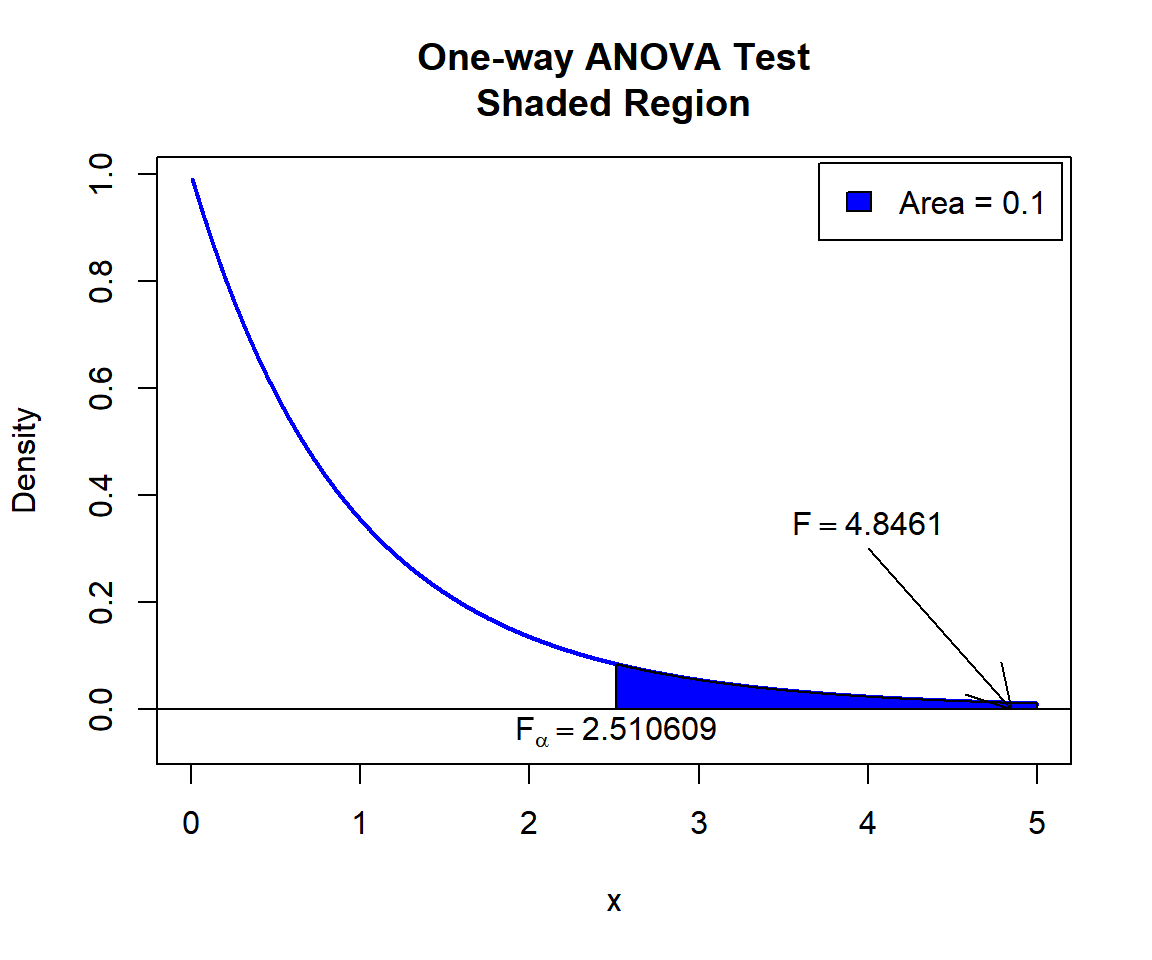
One-way ANOVA Test Shaded Region in R
5 One-way ANOVA Test: Test Statistics, P-value & Degrees of Freedom in R
Here for a one-way ANOVA test, we show how to get the test statistics
(or f-value), sum of squares, mean squares, p-values, and degrees of
freedom from the anova() or oneway.test()
function in R, or by written code.
aov_object = anova(lm(weight ~ feed, data = chickwts))
owt_object = oneway.test(weight ~ feed, data = chickwts,
var.equal = TRUE)
aov_objectAnalysis of Variance Table
Response: weight
Df Sum Sq Mean Sq F value Pr(>F)
feed 5 231129 46226 15.365 5.936e-10 ***
Residuals 65 195556 3009
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
One-way analysis of means
data: weight and feed
F = 15.365, num df = 5, denom df = 65, p-value = 5.936e-10To get the test statistic (or f-value), sum of squares, and mean squares:
F-value
\[\begin{align*} F & = \frac{\left[ \sum_{j=1}^G N_j(\bar y_{j\cdot}-\bar y_{\cdot \cdot})^2 \right] / (G-1)}{\left[ \sum_{j=1}^G \sum_{k=1}^{N_j} (y_{jk}-\bar y_{j\cdot})^2 \right]/ (N-G)}\\ & = \frac{\left[ \sum_{j=1}^G N_j(\bar y_{j.}-\bar y_{\cdot \cdot})^2 \right]/ (G-1)}{\left[ \sum_{j=1}^G (N_j-1) s_j^2 \right]/ (N-G)}. \end{align*}\]
[1] 15.3648 F
15.3648 [1] 15.3648Same as:
# Method 1
y = chickwts$weight; group = chickwts$feed
f_order = order(group); y = y[f_order]; group = group[f_order] #order by group
means = tapply(y, group, mean) #means
lens = tapply(y, group, length) #sizes
G = length(unique(group)); N = length(y) #lengths
ssg = sum(lens*(means-mean(y))^2) #sums of squares
ssr = sum((y-rep(means, lens))^2)
num = ssg/(G-1); denom = ssr/(N-G) #mean squares
F = num/denom
F[1] 15.3648# Method 2
y = chickwts$weight; group = chickwts$feed
means = tapply(y, group, mean) #means
vars = tapply(y, group, var) #variances
lens = tapply(y, group, length) #sizes
G = length(unique(group)); N = length(y) #lengths
ssg = sum(lens*(means-mean(y))^2) #sums of squares
ssr = sum((lens-1)*vars)
num = ssg/(G-1); denom = ssr/(N-G) #mean squares
F = num/denom
F[1] 15.3648Sum of squares and mean squares
[1] 231129.2 195556.0[1] 46225.832 3008.554Same as (based on steps above):
[1] 231129.2 195556.0[1] 46225.832 3008.554To get the p-value:
The p-value is, \(P (F_{df1, df2}>F_{Observed})\)
[1] 5.93642e-10[1] 5.93642e-10Same as:
Note that the p-value depends on the \(\text{test statistics}\) (\(F_{df1, df2} = 15.3648\)), and \(\text{degrees of freedom}\) (5, 65). We
also use the distribution function pf() for the F
distribution in R.
[1] 5.936418e-10To get the degrees of freedom:
The degrees of freedom are \(\text{df1}=5\) and \(\text{df2}=65\).
[1] 5 65 num df denom df
5 65 [1] 5 65Same as:
[1] 5 65The feedback form is a Google form but it does not collect any personal information.
Please click on the link below to go to the Google form.
Thank You!
Go to Feedback Form
Copyright © 2020 - 2026. All Rights Reserved by Stats Codes